What is a mutation?

VARIATION ON A SIMILAR THEME
The consequences of mutations are varied, which is why different terms are used to describe mutations. Among these terms are: variants, genetic variations, polymorphisms (SNP), molecular anomalies or molecular modifications…
As for us, we will be using either one of the three following terms: mutation, genetic variation or molecular anomaly.

HOW MUTATIONS OCCUR
Mutations are a very frequent phenomenon that occur in all organisms and drive the evolution of species.
Mutations can occur spontaneously during cell division, and throughout an organism’s lifetime. Dozens of mutations are believed to occur each time a cell divides, when its DNA is being replicated.
DNA can also be altered following lesions caused by the environment, such as UV radiation, tobacco, asbestos or certain viruses for example.

DIFFERENT TYPES OF MUTATION
DNA can be altered in different ways:
– one nucleotide is replaced by another (point mutation) (for example: acagtg -> acagag);
– one or more nucleotides are deleted or added (for example: acagtg -> acaacgtg);
– DNA sequences of various lengths are found in variable copies either on the same chromosome or on another chromosome. These changes are called chromosomal rearrangements or ‘Copy Number Variations (CNVs)’.
INHERITED AND NON-INHERITED MUTATIONS
Most mutations (notably those found in cancer cells) are somatic, i.e. they occur spontaneously in cells and are not inherited.
If such a mutation does occur in a germ-line cell (eggs or sperm), however, then it will be passed on to the next generation. About 10% of cancers are heritable.

MUTATIONS AND PROTEINS
The majority of mutations known today, which have a clinical impact on cancer, hit regions coding for proteins.
SMALL CAUSES, GREAT EFFECTS
If a mutation hits a gene that codes for a protein, it may affect the protein’s amino-acid sequence and even its 3D structure.
If a protein’s 3D structure is changed, the protein’s function may then be affected bringing about a different phenotype – such as the absence of fingerprints or a disease like cancer.
Certain mutations can cause a given protein to be overexpressed or, on the contrary, not to be expressed at all. Depending on the biological processes the protein is involved in, the phenotype is more or less pronounced.
Several mutations taken together are usually the cause of one phenotype.

MUTATIONS AND CANCER
MUTATIONS THAT CHANGE CELL BEHAVIOUR
The unusual behaviour of cancer cells is caused by an accumulation of mutations in their DNA.
Cancer cells are particularly unstable from a genetic point of view. They carry many mutations and hence proteins whose biological function or expression has been altered – such as those that regulate cell division or the immune response.
Understanding the mutations found in the DNA of the different cells that compose a tumour helps to make a more precise diagnosis and to select, as far as possible, the most appropriate treatment.

HOW TO DETERMINE A MUTATION’S IMPACT ON THE DEVELOPMENT OF CANCER?
MUTATION CATEGORIES
Several hundred mutations can be found in cancer cells. The majority have no important role in the progression of the disease.
Mutations are classified into 3 different categories:
PASSENGER MUTATIONS
Passenger mutations are benign. A priori, they have no direct effect on a tumour’s growth but just seem to accompany pathogenic mutations, either by accident or on an opportunistic basis.
By being able to pinpoint passenger mutations faster, scientists will be able to ignore them and spend more time on other mutations, thus avoiding unnecessary treatments.
DRIVER MUTATIONS
Driver mutations are pathogenic (oncogenic).
They alter a protein’s function and favour the development of cancer.
Identifying driver mutations helps to refine a diagnosis and, in the best of cases, to offer patients treatments that are more adapted.
ACTIONABLE MUTATIONS
Mutations are called ‘actionable’ if they are known to be associated with a specific treatment.
Certain mutations are resistant to treatment. Unveiling them avoids treatments that would be ineffective, as well as their ensuing side-effects.

VALUABLE POINTERS
Below are a few pointers that help scientists interpret mutations:
– if the ‘mutated’ amino acid is also found in a ‘healthy’ population, then it probably has no impact on the protein’s function (passenger mutation);
– if the ‘mutated’ amino acid is also found in other species, then it will probably have no impact on the protein’s function (passenger mutation);
– if the ‘mutated’ amino acid has the same physicochemical properties as the ‘healthy’ amino acid, then there is less chance that it will have an impact on the protein’s function (passenger mutation);
– if the ‘mutated’ amino acid changes the protein’s 3D structure, it may have an impact on the protein’s function or on its interaction with a drug (driver or actionable mutation).
EXAMPLE
Here is an alignment of several amino-acid sequences of the BRAF protein belonging to different species (source www.Swiss-PO.ch).
The region shown is identical – conserved – in every species. This means that it is probably important for proper BRAF function. If there is a change of amino acid in this highly-conserved region then there is a great chance that it will affect BRAF function.
Amino acid V (Valine) in position 600 is conserved in every species (chimpanzee, macaque, mouse, rat, dog, bovine, chicken, zebrafish and frog). When it is replaced by another amino acid such as E (Glutamate) for example, BRAF function is indeed altered.

HOW TO FIND INFORMATION?
Information regarding mutations, when known, is listed in specialized databanks, usually with online open access.
In 2019, the databank COSMIC contained information on more than 6 million mutations found in proteins involved in cancer.

THE NEED TO UPDATE AND SHARE INFORMATION
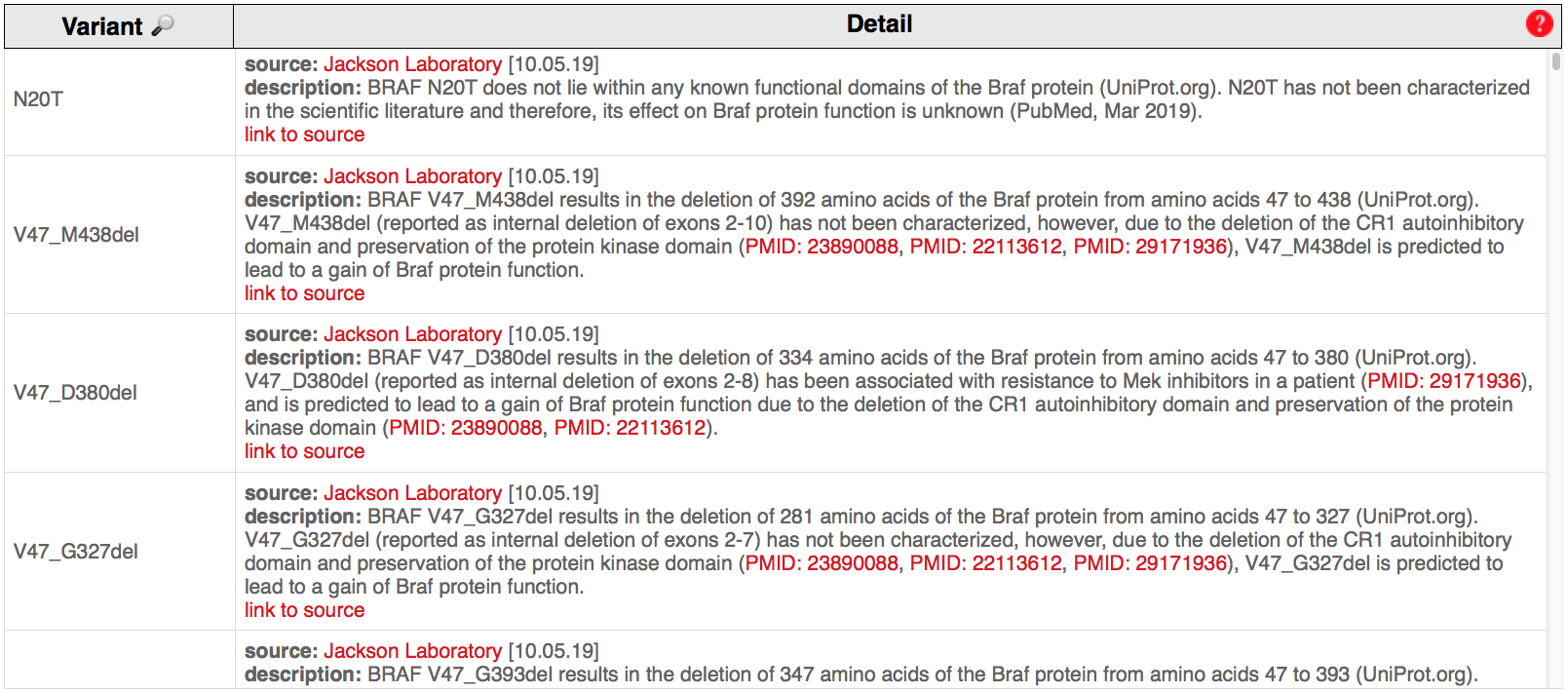
These databanks are regularly updated as they keep up with ongoing research and scientific publications (Example of disclaimer).
The quality and reliability of data are paramount.
Numerous national and international initiatives are currently being undertaken to gather and standardise as much as possible all existing information on the functional and clinical impacts of mutations in cancer and other diseases.

if no information is available ...
In about 25% of cases, a mutation’s effect on a protein’s function – or on the way it affects a given treatment – is either unknown or the data found in databanks are in disagreement.
When this happens, scientists resort to bioinformatics predictive tools – in particular molecular modelling.
Molecular modelling enables scientists to predict and visualize the effect a mutation would have on a protein’s 3D structure. It is thus sometimes possible to predict whether a mutation is liable to have an effect – or not – on an important active site on the protein and/or on the protein’s capacity to interact with a given drug (and inducing drug resistance for instance).

STUBBORN MUTATIONS
Sometimes nothing can be found on the effects certain mutations have.
These mutations are then classified as ‘variants of unknown clinical significance’ (VUS). Classification changes with time, of course, as scientific progress is made.
Each and every one of us carries thousands of VUS but unless we undergo a genetic test, we are unaware of them. When VUS are part of a medical diagnosis however, they do add a measure of uncertainty both for the patient and the doctor.